Radiality Centrality
Definition
Radiality centrality will give high centralities to vertices that are
a short distance to every other vertex in its reachable neighborhood compared to
its diameter.
Radiality out-centrality for a vertex i is computed using the out component gi for the vertex i and is given by (Σj d-dij+1 / (d(n-1)), where dij is the distance from i to j in gi, d is the diameter of gi, and the sum is over the vertices in gi.
Radiality in-centrality for a vertex i is computed using the in component gi for the vertex i and is given by (Σj d-dij+1 / (d(n-1)), where dij is the distance from j to i in gi, d is the diameter of gi, and the sum is over the vertices in gi.
Radiality in-centralities are also known as integration centralities.
The radiality centrality for an isolated vertex is taken to be zero.
[Wolfram Research, Inc., 2014]
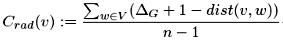 The radiality is a node centrality index. The radiality of a node v is calculated by computing the shortest path between the node v and all other nodes in the graph. The value of each path is then subtracted by the value of the diameter +1 (G+1) and the resulting values are summated. Finally, the obtained value is divided for the number of nodes −1 (n−1). Basically, as the diameter is the maximal possible distance between nodes, subtracting systematically from the diameter the shortest paths between the node v and its neighbors will give high values if the paths are short and low values if the paths are long. Overall, if the radiality is high this means that, with respect to the diameter, the node is generally closer to the other nodes, whereas, if the radiality is low, this means that the node is peripheral. Also here, high and low values are more meaningful when compared to the average radiality of the graph G calculated by averaging the radiality values of all nodes in the graph. As for the closeness, the radiality value should be considered as an average tendency to node proximity or isolation, not definitively informative on the centrality of the individual node. The radiality should be always compared to the closeness and to the eccentricity: a node with high eccentricity + high closeness+ high radiality is a consistent indication of a high central position in the graph.
The radiality is a node centrality index. The radiality of a node v is calculated by computing the shortest path between the node v and all other nodes in the graph. The value of each path is then subtracted by the value of the diameter +1 (G+1) and the resulting values are summated. Finally, the obtained value is divided for the number of nodes −1 (n−1). Basically, as the diameter is the maximal possible distance between nodes, subtracting systematically from the diameter the shortest paths between the node v and its neighbors will give high values if the paths are short and low values if the paths are long. Overall, if the radiality is high this means that, with respect to the diameter, the node is generally closer to the other nodes, whereas, if the radiality is low, this means that the node is peripheral. Also here, high and low values are more meaningful when compared to the average radiality of the graph G calculated by averaging the radiality values of all nodes in the graph. As for the closeness, the radiality value should be considered as an average tendency to node proximity or isolation, not definitively informative on the centrality of the individual node. The radiality should be always compared to the closeness and to the eccentricity: a node with high eccentricity + high closeness+ high radiality is a consistent indication of a high central position in the graph.
In biological terms
The radiality of a node in a biological network, for instance a protein-signaling network, can be interpreted as the probability of a protein to be functionally relevant for several other proteins, but with the possibility to be irrelevant for few other proteins. Thus, a protein with high radiality, compared to the average radiality of the network, will be easily central to the regulation of other proteins but with some proteins not influenced by its activity. Notably, in biological networks could be also of interest to analyze proteins with low radiality, compared to the average radiality of the network, as these proteins, although less relevant for that specific network, are possibly behaving as intersecting boundaries with other networks. Accordingly, a signaling network with a very high average radiality is more likely organizing functional units or modules, whereas a signaling network with very low average radiality will behave more likely as an open cluster of proteins connecting different regulatory modules. All these interpretations should be accompanied to the contemporary evaluation of eccentricity and closeness. [SCARDONI, G.,]
See:
Valente, T.W. and Foreman, R.K., 1998. Integration and radiality: measuring the extent of an individual's connectedness and reachability in a network. Social networks, 20(1), pp.89-105.
Radiality out-centrality for a vertex i is computed using the out component gi for the vertex i and is given by (Σj d-dij+1 / (d(n-1)), where dij is the distance from i to j in gi, d is the diameter of gi, and the sum is over the vertices in gi.
Radiality in-centrality for a vertex i is computed using the in component gi for the vertex i and is given by (Σj d-dij+1 / (d(n-1)), where dij is the distance from j to i in gi, d is the diameter of gi, and the sum is over the vertices in gi.
Radiality in-centralities are also known as integration centralities.
The radiality centrality for an isolated vertex is taken to be zero.
[Wolfram Research, Inc., 2014]
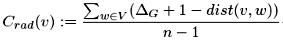
In biological terms
The radiality of a node in a biological network, for instance a protein-signaling network, can be interpreted as the probability of a protein to be functionally relevant for several other proteins, but with the possibility to be irrelevant for few other proteins. Thus, a protein with high radiality, compared to the average radiality of the network, will be easily central to the regulation of other proteins but with some proteins not influenced by its activity. Notably, in biological networks could be also of interest to analyze proteins with low radiality, compared to the average radiality of the network, as these proteins, although less relevant for that specific network, are possibly behaving as intersecting boundaries with other networks. Accordingly, a signaling network with a very high average radiality is more likely organizing functional units or modules, whereas a signaling network with very low average radiality will behave more likely as an open cluster of proteins connecting different regulatory modules. All these interpretations should be accompanied to the contemporary evaluation of eccentricity and closeness. [SCARDONI, G.,]
See:
Valente, T.W. and Foreman, R.K., 1998. Integration and radiality: measuring the extent of an individual's connectedness and reachability in a network. Social networks, 20(1), pp.89-105.
Software
- NetworkAnalyzer
http://med.bioinf.mpi-inf.mpg.de/networkanalyzer/ - CentiBiN
http://centibin.ipk-gatersleben.de/ - CentiLib
http://centilib.ipk-gatersleben.de/ - CentiScaPe
http://www.cbmc.it/~scardonig/centiscape/centiscape.php - Interference
http://www.cbmc.it/~scardonig/interference/Interference.php - MultiNet
http://www.sfu.ca/personal/archives/richards/Multinet/Pages/multinet.htm - Visone
http://visone.info/ - Wolfram
http://www.wolfram.com
References
- Valente, T.W. and Foreman, R.K., 1998. Integration and radiality: Measuring the extent of an individual's connectedness and reachability in a network. Social networks, 20(1), pp.89-105.
- SCARDONI, G., LAUDANNA, C., TOSADORI, G., FABBRI, F. & FAIZAAN, M. CentiScaPe: Network centralities for Cytoscape. http://www.cbmc.it/~scardonig/centiscape/CentiScaPefiles/CentralitiesTutorial.pdf
- Wolfram Research, Inc., Mathematica, Version 10.0, Champaign, IL (2014). http://reference.wolfram.com/language/ref/RadialityCentrality.html