NFC - Neighborhood Functional Centrality
Definition
Neighborhood Functional Centrality (NFC) is a method developed to detect lethal proteins within a protein-protein interaction (PPI) network.
NFC is an integration of the biological function annotation associated with each proteins and the surrounding neighborhood to measure the CENTRALITY of the protein within its active neighborhood.
A protein's lethality correlates more strongly with its “functional centrality" than pure topological centrality.
Functional centrality defined as the topological centrality within a subnetwork of proteins with similar functions.
NFC algorithm consists of two steps. First, for each protein in the interaction graph, construct a local neighborhood graph to compute a nfc score. Then, assess the significance of nfc(u) by computing its corresponding Znfc.
For each vertex u ∈ VPPI, its neighborhood graph is defined as Gu = (Vu,Eu), where:
Relative Specificity Similarity (RSS):
The protein functional similarity between two proteins u and v is defined as: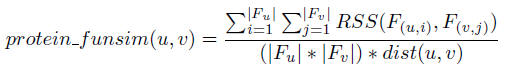 where F(u,i) and F(v,j) denote protein u's i-th and protein v's j-th's functions respectively, and |Fu| denotes the number of functions protein u is annotated with.
where F(u,i) and F(v,j) denote protein u's i-th and protein v's j-th's functions respectively, and |Fu| denotes the number of functions protein u is annotated with.
The neighborhood functional centrality nfc(u) of a protein u is defined as: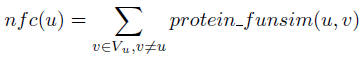 Above defiition quantitates the degree of functional consistency between protein
u and all the other proteins in its neighborhood graph Gu = (Vu,Eu). The value
nfc(u) indicates the functional centrality of protein u in Gu.
Above defiition quantitates the degree of functional consistency between protein
u and all the other proteins in its neighborhood graph Gu = (Vu,Eu). The value
nfc(u) indicates the functional centrality of protein u in Gu.
NFC is an integration of the biological function annotation associated with each proteins and the surrounding neighborhood to measure the CENTRALITY of the protein within its active neighborhood.
A protein's lethality correlates more strongly with its “functional centrality" than pure topological centrality.
Functional centrality defined as the topological centrality within a subnetwork of proteins with similar functions.
NFC algorithm consists of two steps. First, for each protein in the interaction graph, construct a local neighborhood graph to compute a nfc score. Then, assess the significance of nfc(u) by computing its corresponding Znfc.
For each vertex u ∈ VPPI, its neighborhood graph is defined as Gu = (Vu,Eu), where:
Vu = {v | v ∈ VPPI ^ dist(u, v) ≤ 0};
Eu = {(vj, vk) j (vj, vk) ∈ EPPI ^ vj, vk ∈ Vu};
and dist(u, v) is a function that returns the shortest distance between u and v.
Relative Specificity Similarity (RSS):
RSS(termi; termj) = maxDepthGO / (maxDepthGO + γ) . α/ (α + β)
where maxDepthGO is the maximum depth of the GO, α measures the maximum
number of common ancestor terms shared between termi and termj in a single
path, β is the value of the longer distance between termi and termj to their closest
leaf nodes, and γ measures the shortest distance between termi and termj.
The protein functional similarity between two proteins u and v is defined as:
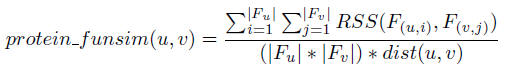
The neighborhood functional centrality nfc(u) of a protein u is defined as:
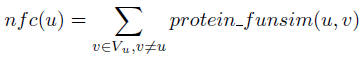
Software
- NFC site
NFC site
References
- TEW, K. L., LI, X.-L. & TAN, S.-H. Functional centrality: detecting lethality of proteins in protein interaction networks. Genome Inform, 2007. World Scientific, 166-177.