Centroid Value
Definition
Centroid value Ccen(v) for node v:
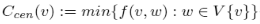 where f(v, w) : = γv(w) - γw(v), and γv(w) is the number of vertex closer to v than to w.
where f(v, w) : = γv(w) - γw(v), and γv(w) is the number of vertex closer to v than to w.
The centroid value is the most complex node centrality index. It is computed by focusing the calculus on couples of nodes (v,w) and systematically counting the nodes that are closer (in term of shortest path) to v or to w. The calculus proceeds by comparing the node distance from other nodes with the distance of all other nodes from the others, such that a high centroid value indicates that a node v is much closer to other nodes. Thus, the centroid value provides a centrality index always weighted with the values of all other nodes in the graph. Indeed, the node with the highest centroid value is also the node with the highest number of neighbors (not only first) if compared with all other nodes. In other terms, a node v with the highest centroid value is the node with the highest number of neighbors separated by the shortest path to v. The centroid value suggests that a specific node has a central position within a graph region characterized by a high density of interacting nodes. Also here, high and low values are more meaningful when compared to the average centrality value of the graph G calculated by averaging the centrality values of all nodes in the graph.
In Biological Terms
The centrality value of a node in a biological network, for instance a protein-signaling network, can be interpreted as the probability of a protein to be functionally capable of organizing discrete protein clusters or modules. Thus, a protein with high centroid value, compared to the average centroid value of the network, will be possibly involved in coordinating the activity of other highly connected proteins, altogether devoted to the regulation of a specific cell activity (for instance, cell adhesion, gene expression, proliferation etc.). Accordingly, a signaling network with a very high average centroid value is more likely organizing functional units or modules, whereas a signaling network with very low average centroid value will behave more likely as an open cluster of proteins connecting different regulatory modules. It can be useful to compare the centroid value to other algorithms detecting dense regions in a graph, indicating protein clusters, such as, for instance, MCODE. [SCARDONI, G.]
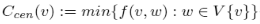
The centroid value is the most complex node centrality index. It is computed by focusing the calculus on couples of nodes (v,w) and systematically counting the nodes that are closer (in term of shortest path) to v or to w. The calculus proceeds by comparing the node distance from other nodes with the distance of all other nodes from the others, such that a high centroid value indicates that a node v is much closer to other nodes. Thus, the centroid value provides a centrality index always weighted with the values of all other nodes in the graph. Indeed, the node with the highest centroid value is also the node with the highest number of neighbors (not only first) if compared with all other nodes. In other terms, a node v with the highest centroid value is the node with the highest number of neighbors separated by the shortest path to v. The centroid value suggests that a specific node has a central position within a graph region characterized by a high density of interacting nodes. Also here, high and low values are more meaningful when compared to the average centrality value of the graph G calculated by averaging the centrality values of all nodes in the graph.
In Biological Terms
The centrality value of a node in a biological network, for instance a protein-signaling network, can be interpreted as the probability of a protein to be functionally capable of organizing discrete protein clusters or modules. Thus, a protein with high centroid value, compared to the average centroid value of the network, will be possibly involved in coordinating the activity of other highly connected proteins, altogether devoted to the regulation of a specific cell activity (for instance, cell adhesion, gene expression, proliferation etc.). Accordingly, a signaling network with a very high average centroid value is more likely organizing functional units or modules, whereas a signaling network with very low average centroid value will behave more likely as an open cluster of proteins connecting different regulatory modules. It can be useful to compare the centroid value to other algorithms detecting dense regions in a graph, indicating protein clusters, such as, for instance, MCODE. [SCARDONI, G.]
Requirements
Require strongly connected network.
Software
- CentiBiN
http://centibin.ipk-gatersleben.de/ - CentiLib
http://centilib.ipk-gatersleben.de/ - CentiScaPe
http://www.cbmc.it/~scardonig/centiscape/centiscape.php - GraphStream
http://graphstream-project.org/ - Interference
http://www.cbmc.it/~scardonig/interference/Interference.php - Web CentiScaPe (FastCent).
http://www.cbmc.it/fastcent/
References
- SCARDONI, G., PETTERLINI, M. & LAUDANNA, C. 2009. Analyzing biological network parameters with CentiScaPe. Bioinformatics, 25, 2857-2859.
DOI: 10.1093/bioinformatics/btp517


- SCARDONI, G., LAUDANNA, C., TOSADORI, G., FABBRI, F. & FAIZAAN, M. CentiScaPe: Network centralities for Cytoscape. http://www.cbmc.it/~scardonig/centiscape/CentiScaPefiles/CentralitiesTutorial.pdf